Publications
Customizing CRISPR–Cas PAM specificity with protein language models
Nature Biotechnology
2026
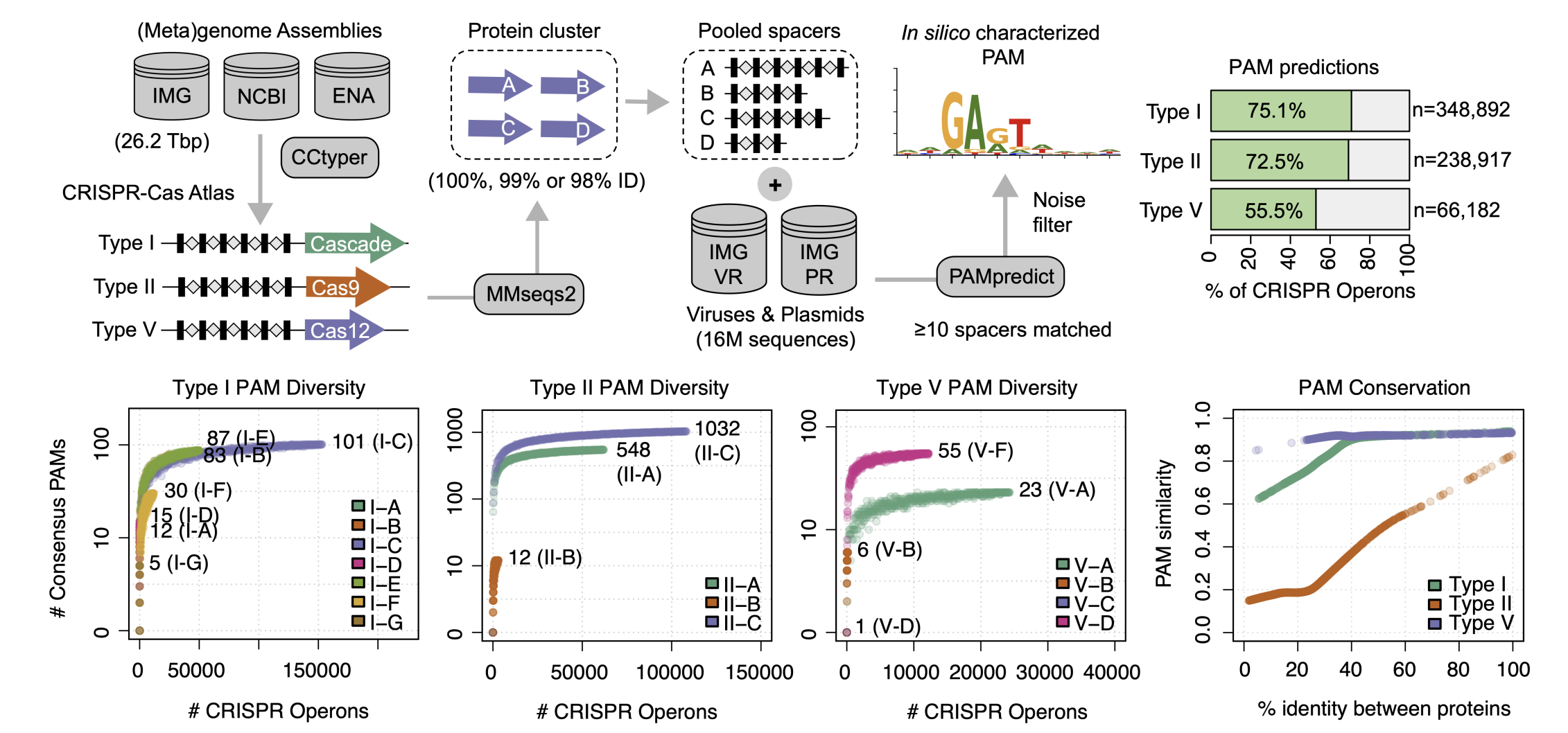
CRISPR–Cas enzymes must recognize a protospacer-adjacent motif (PAM) to edit a genomic site, greatly limiting the range of targetable sequences in a genome. Although engineering strategies to alter PAM specificity exist, they typically require labor-intensive, iterative experimentation. We introduce an evolution-informed deep learning model, Protein2PAM, to efficiently guide the design of Cas protein variants tailored to recognize specific PAMs. Trained on a dataset of over 45,000 CRISPR–Cas PAMs, Protein2PAM rapidly and accurately predicts PAM specificity directly from Cas proteins across type I, II and V CRISPR–Cas systems. Using in silico mutagenesis, the model identifies residues critical for PAM recognition in Cas9 without using structural information. We use Protein2PAM to computationally evolve Nme1Cas9, generating variants with broadened PAM recognition and up to a 50-fold increase in PAM cleavage rates compared to the wild type in vitro. Our machine learning approach allows Cas enzymes to target sequences that were previously inaccessible because of PAM constraints, potentially increasing target flexibility in personalized genome editing.
Access the Paper
- PMID: 39829748
- Biorxiv Preprint
Autoregulation of the MET receptor tyrosine kinase by its intracellular juxtamembrane domain
Biochemical Journal
2025
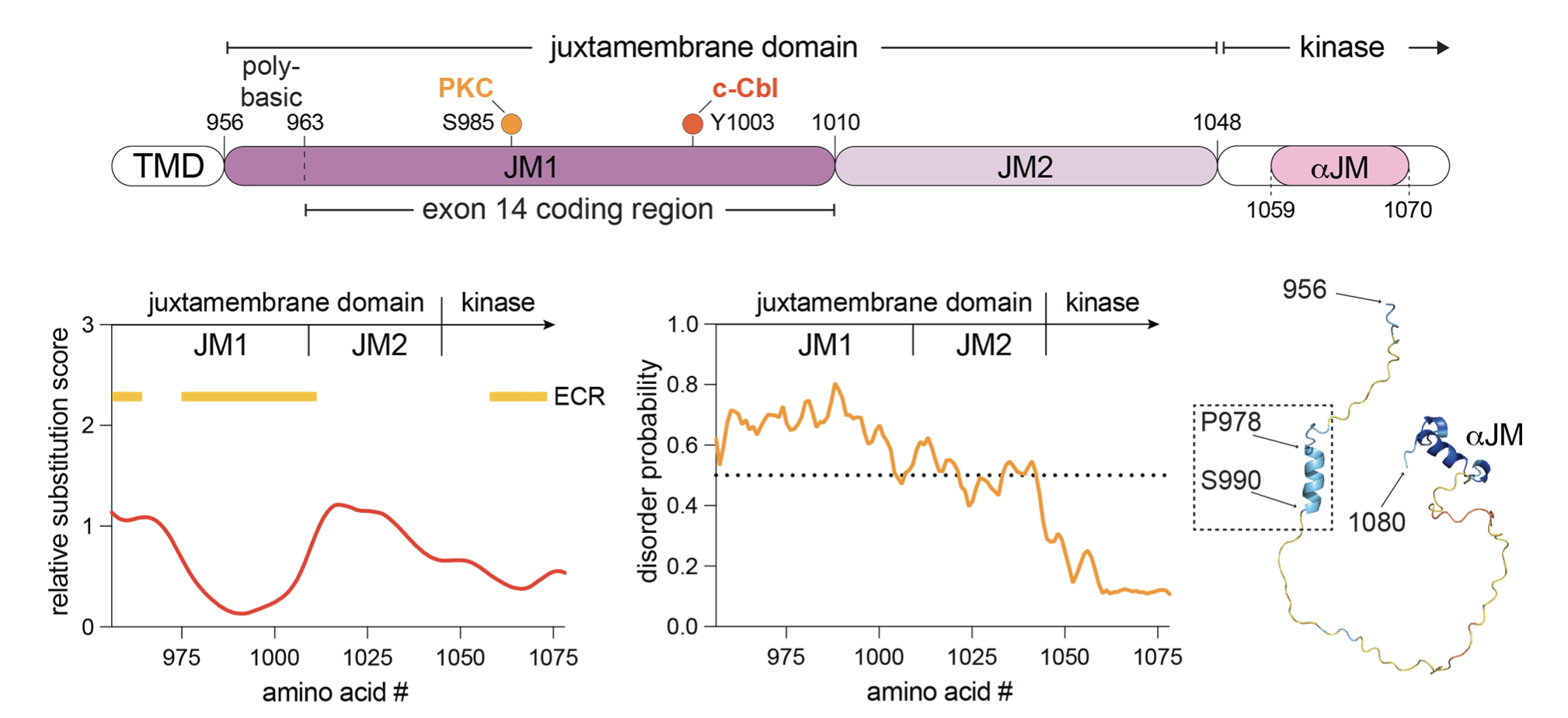
Receptor tyrosine kinases (RTKs) are single-pass transmembrane receptors whose activation is tightly regulated by intra-domain interactions within both their extracellular and intracellular regions. The intracellular juxtamembrane (JM) domain, which links the transmembrane and kinase domains (KDs), often plays a critical role in modulating kinase activity. The MET receptor, activated by hepatocyte growth factor, requires precise regulation to support normal development and wound healing but becomes a potent oncogene when overexpressed or mutated. A common oncogenic lesion in MET, caused by exon 14 skipping, leads to partial deletion of its unusually long intracellular JM domain and is frequently detected in non-small cell lung cancer, as well as pancreatic, liver, and brain cancers. Despite its length and abundance of posttranslational modifications, the functional role of the MET JM domain has remained poorly understood. We have uncovered that this segment regulates the kinetics of MET kinase activation. Specifically, we found that a membrane-proximal, N-terminal region of the JM domain accelerates activation loop phosphorylation, promoting kinase transition to an active state. This regulation does not depend on the oligomeric state of MET but likely acts allosterically to enhance autophosphorylation of the KD. Notably, this function is absent from the closely related MST1R/RON RTK, suggesting it is a unique feature of the MET receptor. Together, these findings uncover a previously unrecognized layer of MET regulation with potential implications for the development of selective therapies targeting MET-driven cancers.
Access the Paper
- PMID: 41383124
- Biorxiv Preprint
Additional Links
MET variants with activating N-lobe mutations identified in hereditary papillary renal cell carcinomas still require ligand stimulation
Molecular Oncology
2025
In hereditary papillary renal cell carcinoma (HPRCC), the hepatocyte growth factor receptor (MET) receptor tyrosine kinase (RTK) mutations recorded to date are located in the kinase domain and lead to constitutive MET activation. This contrasts with MET mutations identified in non-small-cell lung cancer (NSCLC), which lead to exon 14 skipping and deletion of a regulatory domain: In this latter case, the mutated receptor still requires ligand stimulation. Sequencing of MET in samples from 158 HPRCC and 2808 NSCLC patients revealed 10 uncharacterized mutations. Four of these, all found in HPRCC and leading to amino acid substitutions in the N-lobe of the MET kinase, proved able to induce cell transformation, which was further enhanced by hepatocyte growth factor (HGF) stimulation: His1086Leu, Ile1102Thr, Leu1130Ser, and Cis1125Gly. Similar to the variant resulting in MET exon 14 skipping, the two N-lobe MET variants His1086Leu and Ile1102Thr were found to require stimulation by HGF in order to strongly activate downstream signaling pathways and epithelial cell motility. The Ile1102Thr mutation also displayed transforming potential, promoting tumor growth in a xenograft model. In addition, the N-lobe-mutated MET variants were found to trigger a common HGF-stimulation-dependent transcriptional program, consistent with an observed increase in cell motility and invasion. Altogether, this functional characterization revealed that N-lobe variants still require ligand stimulation, in contrast to other RTK variants. This suggests that HGF expression in the tumor microenvironment is important for tumor growth. The sensitivity of these variants to MET inhibitors opens the way for use of targeted therapies for patients harboring the corresponding mutations.
Additional Links
Mapping kinase domain resistance mechanisms for the MET receptor tyrosine kinase via deep mutational scanning
eLife
2025
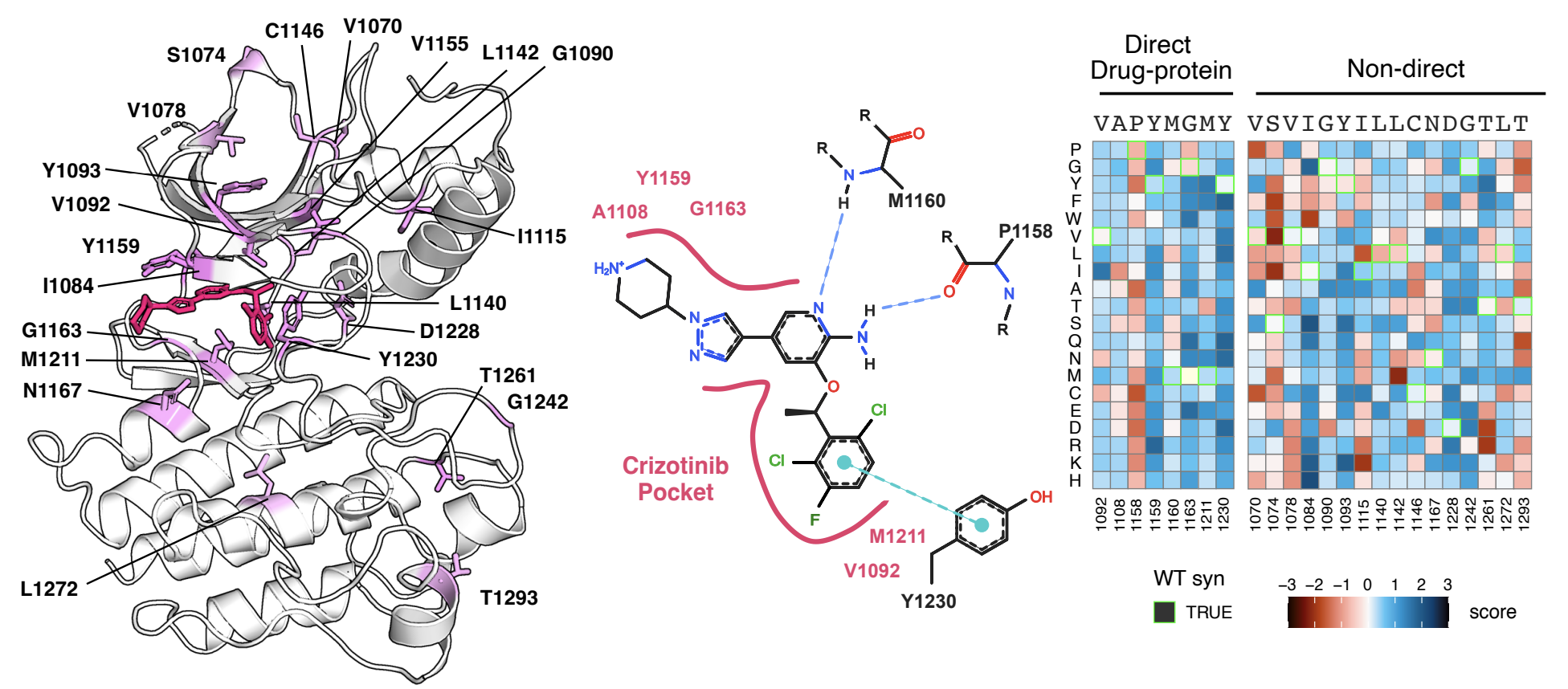
Mutations in the kinase and juxtamembrane domains of the MET Receptor Tyrosine Kinase are responsible for oncogenesis in various cancers and can drive resistance to MET-directed treatments. Determining the most effective inhibitor for each mutational profile is a major challenge for MET-driven cancer treatment in precision medicine. Here, we used a deep mutational scan (DMS) of ~5764 MET kinase domain variants to profile the growth of each mutation against a panel of 11 inhibitors that are reported to target the MET kinase domain. We validate previously identified resistance mutations, pinpoint common resistance sites across type I, type II, and type I ½ inhibitors, unveil unique resistance and sensitizing mutations for each inhibitor, and verify non-cross-resistant sensitivities for type I and type II inhibitor pairs. We augment a protein language model with biophysical and chemical features to improve the predictive performance for inhibitor-treated datasets. Together, our study demonstrates a pooled experimental pipeline for identifying resistance mutations, provides a reference dictionary for mutations that are sensitized to specific therapies, and offers insights for future drug development.
Access the Paper
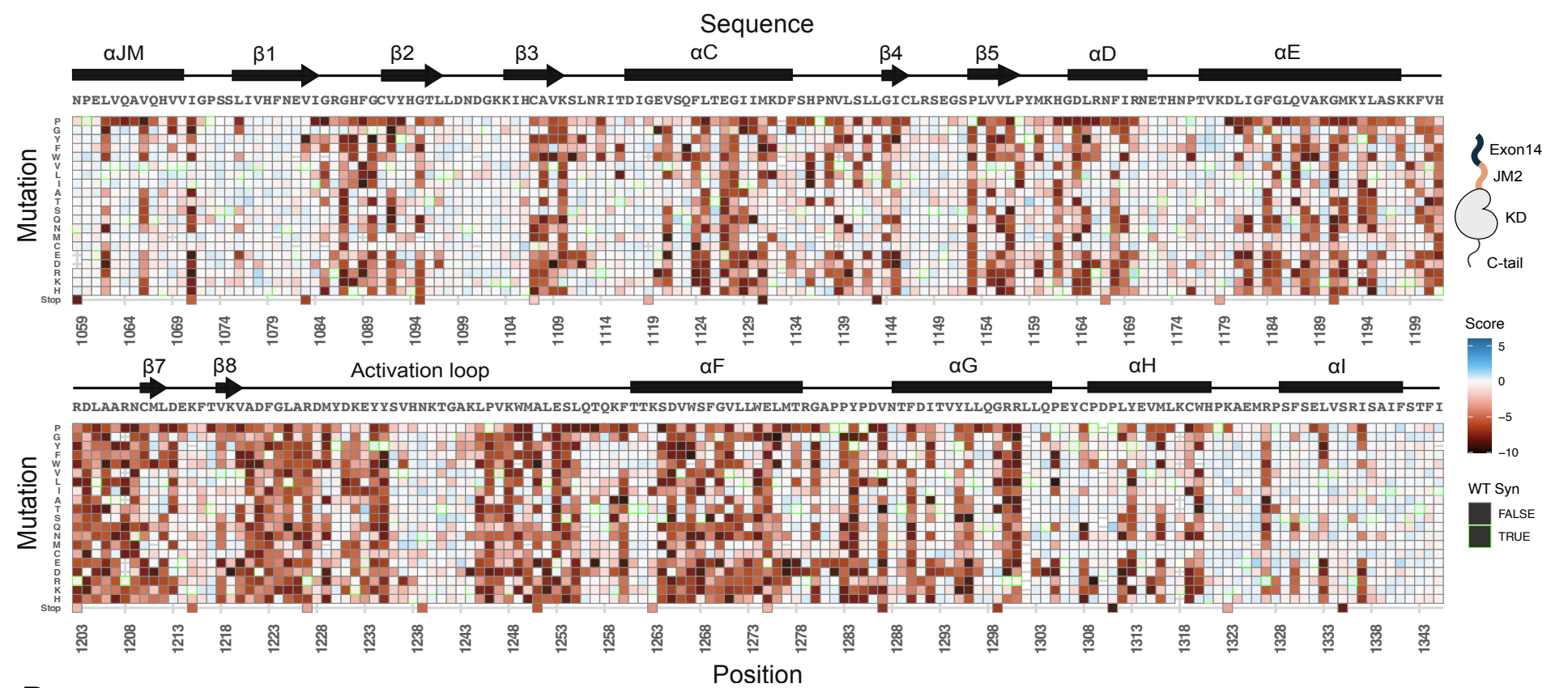
Conserved regulatory motifs in the juxtamembrane domain and kinase N-lobe revealed through deep mutational scanning of the MET receptor tyrosine kinase domain
eLife
2024
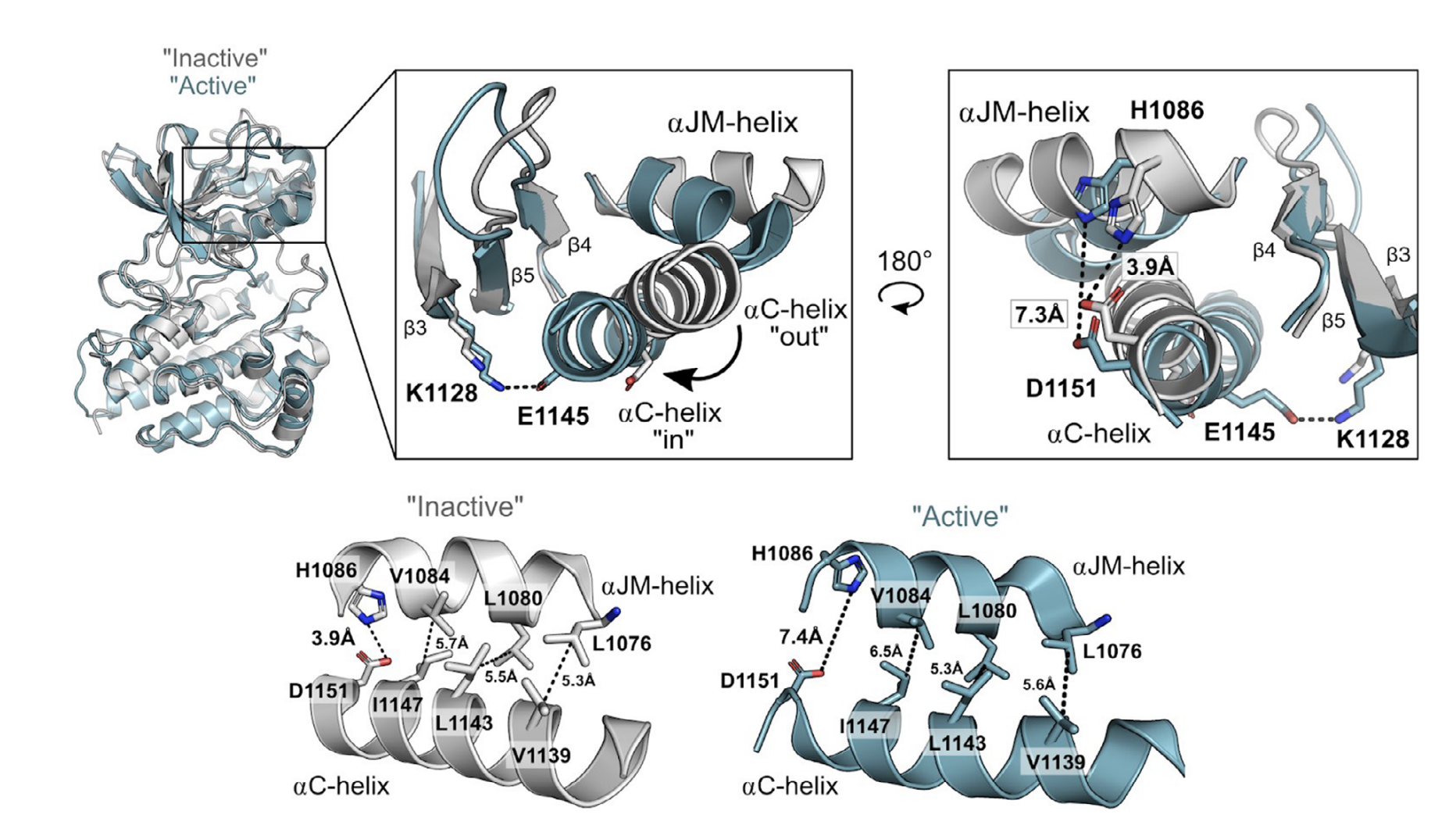
MET is a receptor tyrosine kinase (RTK) responsible for initiating signaling pathways involved in development and wound repair. MET activation relies on ligand binding to the extracellular receptor, which prompts dimerization, intracellular phosphorylation, and recruitment of associated signaling proteins. Mutations, which are predominantly observed clinically in the intracellular juxtamembrane and kinase domains, can disrupt typical MET regulatory mechanisms. Understanding how juxtamembrane variants, such as exon 14 skipping (METΔEx14), and rare kinase domain mutations can increase signaling, often leading to cancer, remains a challenge. Here, we perform a parallel deep mutational scan (DMS) of the MET intracellular kinase domain in two fusion protein backgrounds: wild-type and METΔEx14. Our comparative approach has revealed a critical hydrophobic interaction between a juxtamembrane segment and the kinase ⍺C-helix, pointing to potential differences in regulatory mechanisms between MET and other RTKs. Additionally, we have uncovered a β5 motif that acts as a structural pivot for the kinase domain in MET and other TAM family of kinases. We also describe a number of previously unknown activating mutations, aiding the effort to annotate driver, passenger, and drug resistance mutations in the MET kinase domain.
Access the Paper
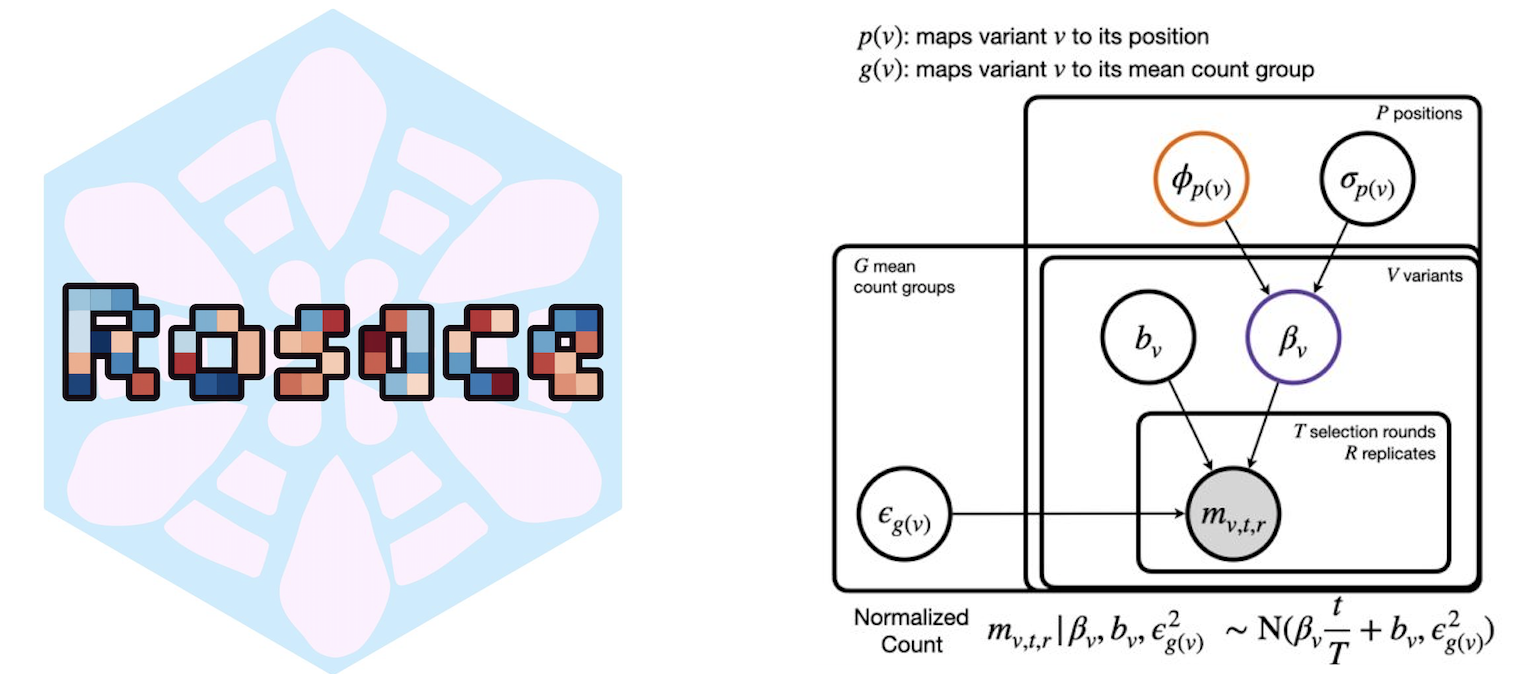
Rosace: a robust deep mutational scanning analysis framework employing position and mean-variance shrinkage
Genome Biology
2024
Deep mutational scanning (DMS) measures the effects of thousands of genetic variants in a protein simultaneously. The small sample size renders classical statistical methods ineffective. For example, p-values cannot be correctly calibrated when treating variants independently. We propose Rosace, a Bayesian framework for analyzing growth-based DMS data. Rosace leverages amino acid position information to increase power and control the false discovery rate by sharing information across parameters via shrinkage. We also developed Rosette for simulating the distributional properties of DMS. We show that Rosace is robust to the violation of model assumptions and is more powerful than existing tools.
Access the Paper
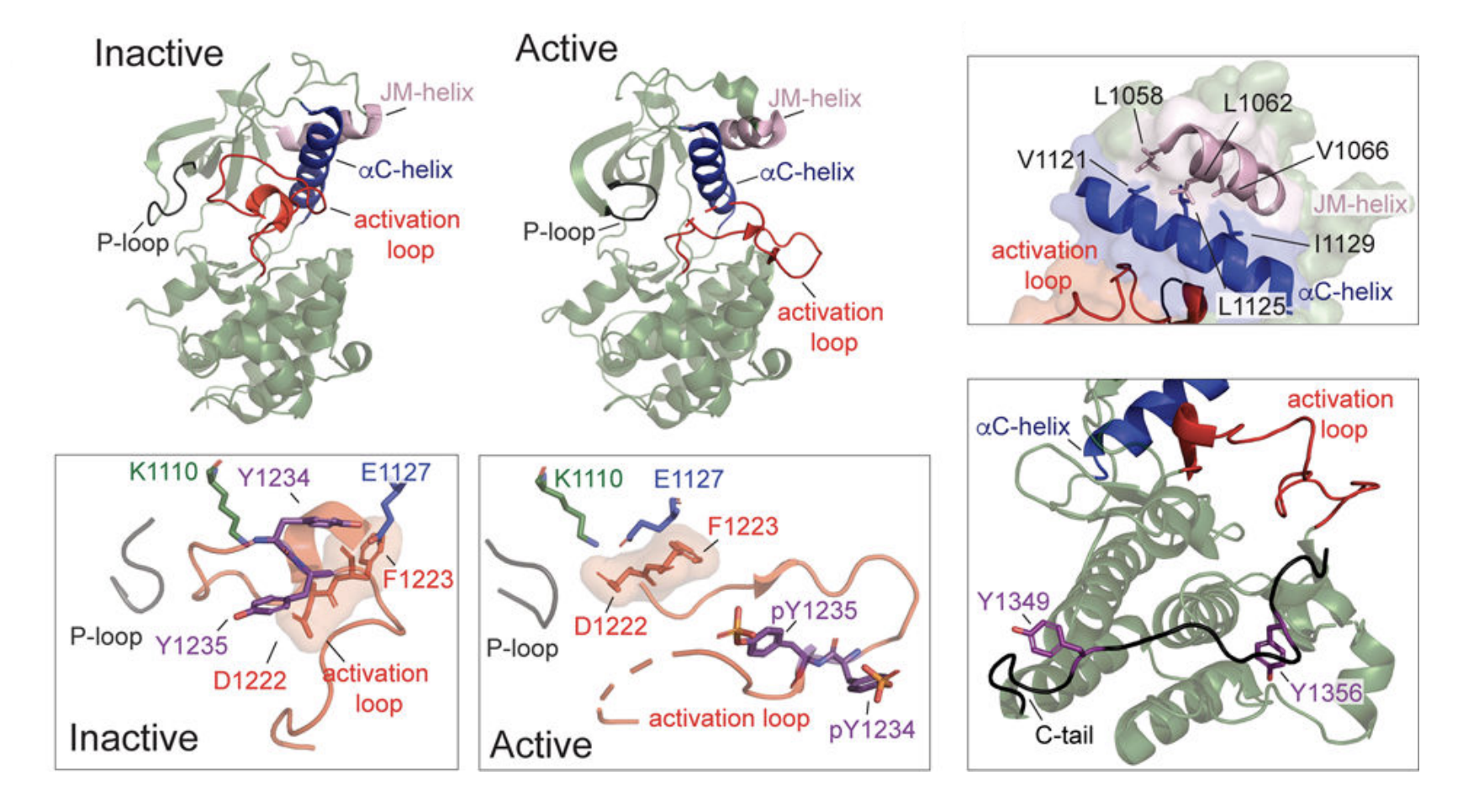
State of the structure address on MET receptor activation by HGF
Biochemical Society Transactions
2021
The MET receptor tyrosine kinase (RTK) and its cognate ligand hepatocyte growth factor (HGF) comprise a signaling axis essential for development, wound healing and tissue homeostasis. Aberrant HGF/MET signaling is a driver of many cancers and contributes to drug resistance to several approved therapeutics targeting other RTKs, making MET itself an important drug target. In RTKs, homeostatic receptor signaling is dependent on autoinhibition in the absence of ligand binding and orchestrated set of conformational changes induced by ligand-mediated receptor dimerization that result in activation of the intracellular kinase domains. A fundamental understanding of these mechanisms in the MET receptor remains incomplete, despite decades of research. This is due in part to the complex structure of the HGF ligand, which remains unknown in its full-length form, and a lack of high-resolution structures of the complete MET extracellular portion in an apo or ligand-bound state. A current view of HGF-dependent MET activation has evolved from biochemical and structural studies of HGF and MET fragments and here we review what these findings have thus far revealed.

